The next generation of LRC-TriCEPS
Now available
TriCEPS v.3.0 enables receptor identification for your orphan ligands at the surface of living cells without genetic manipulation. Key features of the new TriCEPS v.3.0 and LRC- TriCEPS technology are:
- Reduction of the number of cells needed:
- TRICEPS v.3.0 LRC experiments require 5-10 fold less starting material for successful receptor identification compared to TRICEPS v.2.0.
- High coverage of the surfaceome.
- A modified LRC workflow enabled by TriCEPS v.3.0 allows for the theoretical identification of up to 85% of all putative cell surface proteins.
- Identification of N-, C-, O- glycosylated targets
More sensitive identification of low copy number cell surface receptors through whole protein pull-down.
Contact form
Please fill out all mandatory (*) fields.
Decoding the extracellular interactome using LRC-TriCEPS
Ligand-receptor capture
The LRC-TriCEPS technology was invented by Andreas Frei and Bernd Wollscheid from ETH Zurich who published the technology in Nature Biotechnology Nature Biotechnology (Frei et al.(2012) Nature Biotechnology 30:997-1001) and further developed it to HATRIC-LRC (Sobotzki et al. Nature Communications, volume 9, Article number: 1519 (2018) doi:10.1038/s41467-018-03936-z)
Orphan ligands:
- Extracellular proteins
- Peptide ligands
- Antibodies
- Viruses
- Engineered affinity binders
Dualsystems Biotech AG is a biotech company providing analytic services for protein-protein interactions.
If you are looking for the targets of your ligand – peptide, protein, antibody, and virus – that are relevant in the living organism, call us on +41 44 738 50 00 or fill in the form on the right hand. We will contact you to see how LRC-TriCEPS technology (ligand-receptor capture) can answer your questions.
LRC-TriCEPS is a novel approach enabling the identification of cell surface receptors and off-targets on living cells for a wide range of orphan ligands. Advantages of LRC-TriCEPS compared to other approaches:
- No genetic manipulation
- No hypothesis needed
- Targets are embedded in plasma membrane in their natural microenvironment
- Cells are alive during interaction
Availability
The TriCEPS-based ligand-receptor capture technology is now available at Dualsystems.
LRC-TriCEPS is a novel approach enabling the identification of cell surface receptors and off-targets on living cells for a wide range of orphan ligands, such as:
- Peptides
- Protein
- Antibodies
- Engineered affinity binders
- Viruses
Availability
The TriCEPS-based ligand-receptor capture technology is now available.
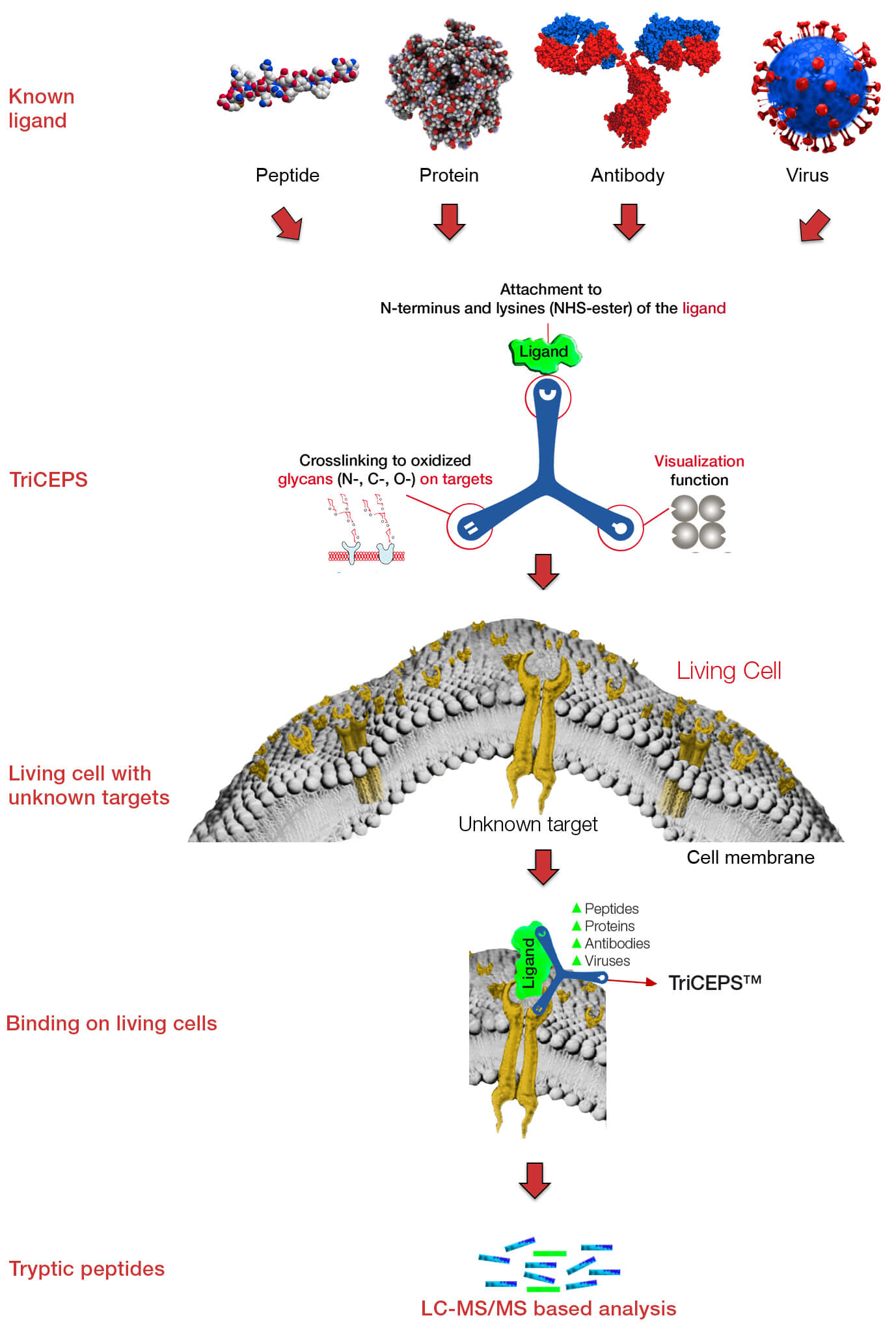
Principle of LRC-TriCEPS
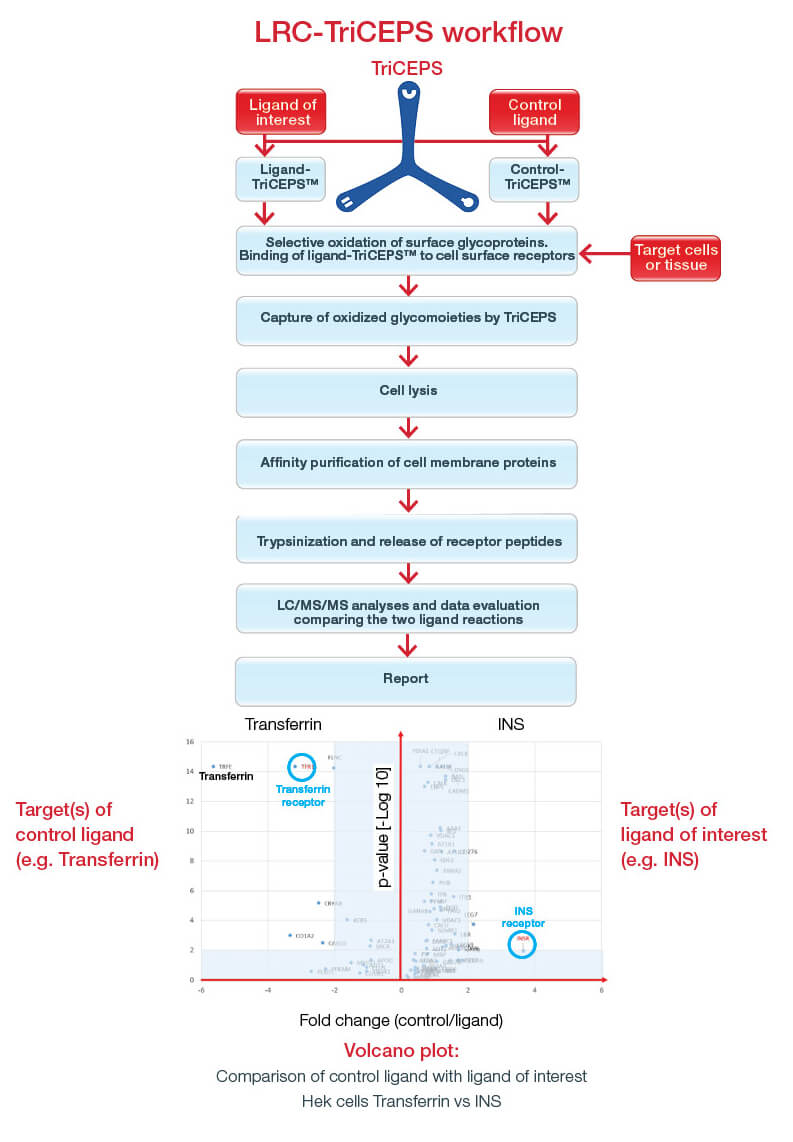
LRC-TriCEPS is a fast and sensitive approach to discover cell surface N-, C-, and O-glycosylated receptors for a ligand of interest.
- Label ligand with the TriCEPS chemoproteomic reagent
- Activate target cells or native tissue to oxidize cell surface glycans
- Incubate ligand-TriCEPS complex with activated cells
- TriCEPS immediately links ligand and cognate receptor(s)
- Isolate all membrane proteins with the support of TriCEPS
- Tryptic digest of all membrane proteins
- Identification of peptides by mass spectrometry
- Analysis of data using statistics and bioinformatics
LRC-TriCEPS – a trifunctional chemoproteomics reagent
TriCEPS structure:
- NHS-ester for attachment to the ligand
- Protected hydrazine function for capturing the interacting receptor
- Pull out function for purification of ligand-receptor peptides
Requirements
Requirements for one LRC experiment: Ligand in triplicate compared to control in triplicate
- 300 µg ligand containing a free amino group
- 60 Mio cells or 50 µm tissue slice
The LRC-TriCEPS platform
enables the identification of protein targets at the cell surface of living cells of ligands such as peptides, proteins, and viruses.
Why is the LRC-TriCEPS platform the best way to identify surface protein interactions or do screenings for peptides, antibodies, or other biologics?
The experiment is done on living cells, without the need to genetically modify the cells or have a hypothesis about the likely target. It can be applied to any mammalian cell type (primary cells or cell lines). During the interaction of your ligand of interest (peptide, protein, virus, or similar) with its unknown target, the cells are alive and can be brought into the desired state (e.g., activated T-cells or differentiated cells), providing a more accurate reflection of what will later occur in the human body, specifically in patients.
Targets and Off-targets, Mode of Action studies for biologics screening or protein interaction mapping
With the LRC-TriCEPS platform, cell surface protein targets and off-targets of biologics (peptides, proteins, antibodies, antibody-drug conjugates (ADC), virus-like particles (VLPs), viruses, protein-containing vesicles, etc.) can be identified. This helps to understand the mode of action (MoA) of biologics and supports the selection of the best biologics candidate by identifying off-targets and comparing several drug candidates in one experiment.
By using different cell types representing various organs of the human body, the LRC-TriCEPS platform can also test whether different biologics drug candidates bind to different proteins on the cell surface of different cell types.
Novel Drug Targets to Broaden Your Drug Pipeline and Find New Ways to Treat Diseases
The LRC-TriCEPS platform is useful for identifying unknown protein-ligand interactions at the cell surface to discover novel drug targets and modes of action by pinpointing endogenous protein interactions. Understanding which proteins are involved at the cell surface is key to discovering novel drug targets.
Case Studies
Immune Oncology Novel Drug Target Discovery
- Challenge: What is the target of the checkpoint inhibitor Vista?
- Background: Vista is a receptor expressed on cancer cells that inhibits activated T-cells from recognizing and killing cancer cells. The LRC-TriCEPS experiment used the extracellular domain of Vista fused with an Fc-tag as the ligand to identify the target on activated T-cells from different donors. At physiological pH 7.4, no target could be identified, but at acidic pH 6.0, PSGL-1 was discovered as the target. The acidic dependency of this interaction was confirmed using different techniques. This experiment mimicked the acidic tumor environment in patients, leading to the identification of a novel drug target.
- Take-home message:
- Novel drug target PSGL-1 identified using the LRC-TriCEPS platform.
- The microenvironment is essential for receptor-ligand interactions.
- Receptor discoveries can best be studied on living cells for a more accurate reflection of patient responses.
- Link to publication: Vista Interaction with PSGL-1 Published in Nature
Novel Drug Target for Asthma Treatment
- Challenge: What proteins are targeted at the cell surface by house dust mite, leading to allergic reactions?
- Background: Researchers at the University of Central Florida wanted to understand how house dust mite induces allergies that may lead to asthma. A house dust mite protein extract was used as a ligand in an LRC-TriCEPS experiment on dendritic cells, identifying LMAN1 as the target. The group confirmed that LMAN1 is downregulated in asthmatic patients and could potentially be used as a novel drug target.
- Take-home message:
- The LRC-TriCEPS platform can identify targets of complex protein or peptide mixtures on the surface of living cells.
- Link to publication: Identification of a Receptor for House Dust Mite Allergens
Mode of Action of a Cytokine Inhibitor (TIMP-1)
- Challenge: What is the mode of action of the cytokine inhibitor TIMP-1?
- Background: TIMP-1 plays a role in inflammatory diseases and cancer, but its interaction with immune cells was unclear. The LRC-TriCEPS platform identified a novel target on the surface of a monocyte cell line, showing that TIMP-1 triggers monocyte activation, a key mechanism in diseases like cancer, sepsis, and acute pancreatitis.
- Take-home message:
- The LRC-TriCEPS platform identified a novel mechanism of action by revealing that TIMP-1 activates monocytes, a crucial factor in inflammatory diseases.
Link to publication: TIMP-1 is a Novel Ligand of Amyloid Precursor Protein and Triggers a flammatory Phenotype in Human Monocytes
Cardiac-Specific Peptide Targeting
- Challenge: How does a cardiac-specific peptide enter cardiac cells?
- Background: Before the LRC-TriCEPS screen, there was no hypothesis regarding the target or how the peptide entered cardiac cells. The screen revealed that a voltage-gated potassium channel was the entry point for the 12-amino acid peptide.
- Take-home message:
- The target of a peptide or protein screen may not always be a receptor; in this case, it was a channel.
- Link to publication: Cardiac Targeting Peptide: A Novel Cardiac Vector Studies in Bio-Distribution, Imaging Application, and Mechanism of Transduction
Customers Testimonials – LRC-TriCEPS Service
Testimonials from our customers who have used the LRC-TriCEPS technology – in collaboration with Dualsystems Biotech AG.
LRC-TriCEPS customers worldwide
Over 200 satisfied customers from 28 countries.
We identify targets / off-targets of your ligand
If you want to get support for your project
LRC-TriCEPS / HATRIC-LRC Publications
Concerning the LRC-TriCEPS or HATRIC-LRC platforms.














 Dr Patric Delhanty
Dr Patric Delhanty



 (Quim) Madrenas, MD, PhD, FCAHS
(Quim) Madrenas, MD, PhD, FCAHS





 De'Broski R. Herbert Ph.D.
De'Broski R. Herbert Ph.D.