HATRIC-based identification of receptors for orphan ligands (small molecules)
HATRIC-LRC (Ligand Receptor Capture) enables now the identification of small molecule targets on the surface of living cells! Even primary cells!
In their latest publication, the Wollscheid group (ETH Zurich, Switzerland) demonstrated how HATRIC-LRC can be used to identify orphan ligand targets from as few as 1 million cells, enabling the LRC technology to be performed on rare primary cells or clinical specimens. What is more, next to de-orphaning proteins, viruses and antibodies, HATRIC-LRC can now be used to identify the unknown targets of small molecules on the cell membrane of living cells in order to elucidate the mode of action of the molecule of interest.
Nadine Sobotzki, Michael A. Schafroth, Alina Rudnicka, Anika Koetemann, Florian Marty, Sandra Goetze, Yohei Yamauchi, Erick M. Carreira & Bernd Wollscheid
Nature Communications, volume 9, Article number: 1519 (2018) doi:10.1038/s41467-018-03936-z,
Published online:
Abstract
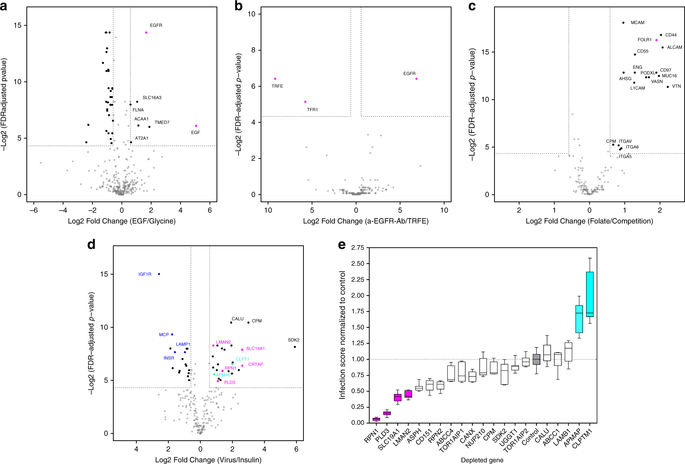
Cellular responses depend on the interactions of extracellular ligands, such as nutrients, growth factors, or drugs, with specific cell-surface receptors. The sensitivity of these interactions to non-physiological conditions, however, makes them challenging to study using in vitro assays. Here we present HATRIC-based ligand receptor capture (HATRIC-LRC), a chemoproteomic technology that successfully identifies target receptors for orphan ligands on living cells ranging from small molecules to intact viruses. HATRIC-LRC combines a click chemistry-based, protein-centric workflow with a water-soluble catalyst to capture ligand-receptor interactions at physiological pH from as few as 1 million cells. We show HATRIC-LRC utility for general antibody target validation within the native nanoscale organization of the surfaceome, as well as receptor identification for a small molecule ligand. HATRIC-LRC further enables the identification of complex extracellular interactomes, such as the host receptor panel for influenza A virus (IAV), the causative agent of the common flu.